High Performance Magnetic Nanoparticle (FG beads)
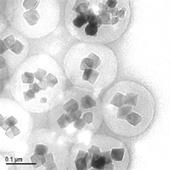

In pharmaceutical research, it is extremely important to perform isolate and identify the protein that is the target of a drug (compound) in vivo. However in the past, isolation and identification of the drug target protein were extremely difficult, and were a problem which required large amounts of time and effort. In order to overcome this problem, through joint research with Professor Hiroshi Handa at the Tokyo Institute of Technology, Tamagawa Seiki has developed the new nanomagnetic particles "FG beads" and the automated screening system "Target Angler".
FG beads are approximately 200 nm in diameter that are composed of multiple ferrite particles coated with a special polymer called poly-GMA (glycidyl methacrylate). The FG beads that are manufactured using this original technology are used as materices for affinity purification and provide characteristics that are superior to conventional materices, allowing one-step purification of the drug target protein. The development of an automated screening system that magnetically separates and disperses the FG beads makes it possible to automate the affinity purification process, process multiple samples simultaneously, and shorten the time required.
Features
- Extremely low non-specific binding:
poly-GMA (Glycidyl methacrylate) coated magnetic nanoparticles - Excellent recovery of target proteins:
200 nm particles have a large surface area and a high dispersibility - High stability in organic solvents:
various compounds can be immobilized on the beads
Product Information
Various Magnetic Beads
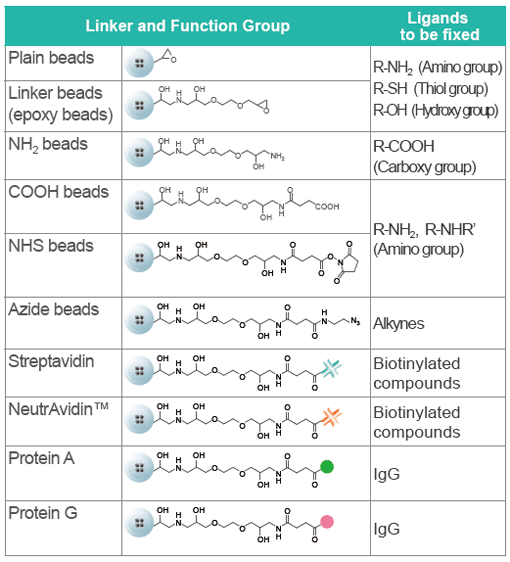
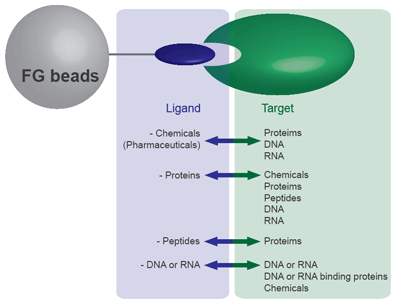
FG beads can capture a variety of substances, including chemicals (drugs), proteins, and DNA. From the 9 types of FG beads, select the type with the optimal surface modification according to the functional group of the substance you want to bind. Because FG beads are resistant to various organic solvents, they are able to bind a variety of ligands. (Avoid using streptavidin beads or other protein-binding FG beads in organic solvents.) The beads with ligands immobilized can be used for affinity purification of the target biological substance.
Linkers and Functional Groups
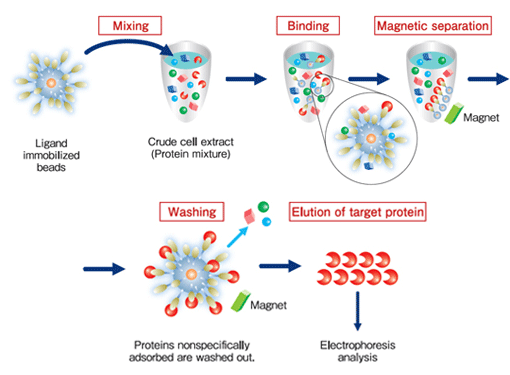
Purification process
Protein purification (affinity purification) using FG beads is performed in 3 stages:
binding, washing, and elution.
During the binding process, the beads immobilized ligands are mixed with the crude cell extract, and the proteins with affinity for the ligands bind to the beads. During the washing process, the proteins that have bound non-specifically to beads (not the proteins bound specifically to the ligands) are washed off. During the elution process, the specifically-bound proteins are separated from the ligands and recovered. At each of these processes, magnets are used to separate immobilized beads from the crude cell extract, washing buffer, or elution buffer.
Comparison with other magnetic beads

Affinity purification of MTX binding proteins Immobilization of MTX on commercial magnetic beads was done in the same manner as in the case of FG beads. |
 |
Applications
Application Note (PDF)
| Application Note 1: | Isolation of drug target protein |
|---|---|
| Application Note 2: | Cell separation, THP-1 |
| Application Note 3: | Isolation of drug target protein Protein Kinase inhibitors (Bisindolylmareimide) |
| Application Note 4: | Isolation of drug target protein Histone Deacetylase Inhibitor (Vorinostat) |
| Application Note 5: | Immunoprecipitation |
Application Examples
Purification of target protein of Thalidomide (elucidation of the teratogenic mechanism)
CRBN (Cereblon) and DDB1 were isolated from human cell extract using thalidomide fixed beads. As a result, the teratogenic mechanism of thalidomide was elucidated.
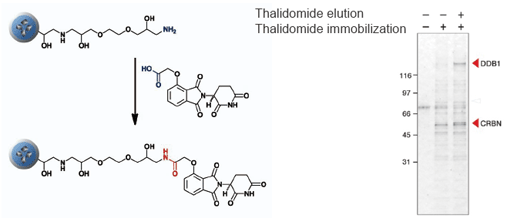
T. Ito et al., Science 327 (2010) 1345
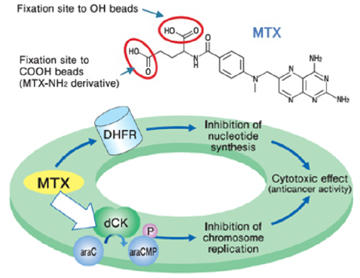
Purification of novel target protein of MTX (methotrexate)
When MTX was fixed via different site, a novel protein was purified and identified as deoxycytidine kinase (dCK). As a result, a possible mechanism of synergistic effect between MTX and ara-C on malignant lymphoma was proposed.
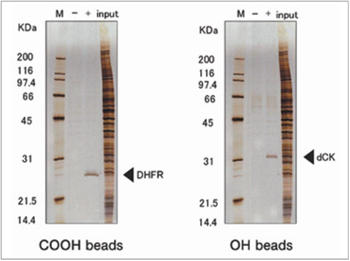
H. Uga et al., Mol. Pharmacol. 70 (2006) 1832
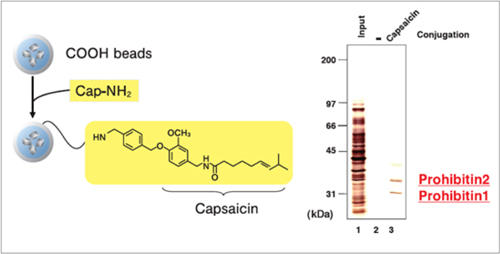
Purification of target proteins of Capsaicin
Prohibitin 1 and prohibitin 2 were isolated from human myeloid leukemia NB4 cell extract using capsaicin derivative (Cap-NH2) fixed beads. As a result, the apoptosis induction mechanism of capsaicin was elucidated.
Elucidation of mechanism of enteropathogenic E. coli enfection
EspB is a protein of enteropathogenic E. coli (EPEC) essential for infection in humans, Myosin was isolated from human cell extract using EspB fixed beads. As a result, the mechanism of EPEC infection was elucidated.
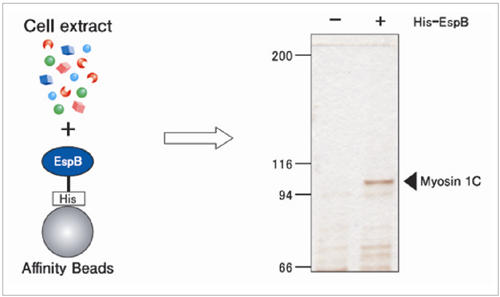

Y. lizumi et al., Cell Host & Microbe. 2 (2007) 383
Protocols
Protocol for Specific Application
Chemical pulldown
| E001 | Screening by using ligand immobilized beads | 259KB |
|---|---|---|
| E003 | Immobilization of ligands (compounds with phenol groups or NH2 groups) on epoxy beads | 111KB |
| E004 | Immobilization of ligands (carboxylic compounds) on OH beads | 78KB |
| E005 | Immobilization of ligands (carboxylic compounds) on NH2 beads | 97KB |
| E008 | Immobilization of ligands (compounds with NH2 groups) on COOH beads | 139KB |
| E012 | Competitive inhibition | 261KB |
| E013 | Drug elution | 260KB |
| E014 | Immobilization of ligands (compounds with NH2 groups) on NHS beads | 75KB |
| E015 | Immobilization of MTX aminated derivatives on NHS beads | 66KB |
| E108 | Immobilization of Ligand on Streptavidin beads | 102KB |
| E109 | Immobilization of ligands (alkyne structure compounds) on azide beads using click chemistry reaction | 65KB |
| E201 | Quantifying the amount of ligand immobilization by HPLC (High Performance Liquid Chromatography) | 1,329KB |
Immunoprecipitation, Protein-protein interaction
| E011 | Immunoprecipitation | 77KB |
|---|---|---|
| E101 | Immobilization of proteins on COOH beads | 83KB |
| E102 | Immobilization of His-Tag proteins on Ts beads | 89KB |
| E105 | Immobilization of Antibodies or Proteins on NHS beads | 23KB |
| E106 | Immobilization of Antibodies or Proteins on Epoxy Beads | 65KB |
| E107 | Direct Quantification of Immobilized Proteins (Antibodies) | 107KB |
| E108 | Immobilization of Ligand on Streptavidin beads | 102KB |
| E110 | Immobilization of antibodies on Protein A beads and Protein G beads | 103KB |
Purification of DNA and RNA binding substances
| E001 | Screening by using ligand immobilized beads | 259KB |
|---|---|---|
| E301 | Immobilization of double strand DNA on Plain beads | 143KB |
Tips; Background Reduction
Immobilization
- Perform centrifugal separation.
The nano size of FG beads gives them excellent dispersibility. As a result, magnetic separation in an organic solvent may be difficult, and it is necessary to recover the FG beads by centrifugal separation. - Disperse the beads well.
Centrifugal separation causes the FG beads to agglutinate strongly, making it difficult to disperse them. Ordinarily a manual dispersion method or ultrasound would be used to disperse the beads. Although ultrasound separates the beads easily, caution is required due to the possibility of damaging the proteins. Therefore in general it is recommended that ultrasound be used when binding low molecular weight compounds, and that manual dispersion be used when binding proteins. The manual method involves resting the bottom of the micro tube in a plastic test tube stand and moving it roughly to disperse the beads. Depending on the type of micro tube, when using the manual method to disperse the beads, the bottom of the tube may crack or leakage from the lid may occur. Use a micro tube that is strong and has a lid with a tight fit. We recommend the use of cap locks.
Dispersion by sonication
Screening
- Perform magnetic separation
When centrifugal separation is performed, heavy proteins and insoluble proteins are also precipitated, raising the level of the background. Because FG beads have high dispersibility, magnetic separation requires time (in some cases 5 minutes or longer). However using magnetic separation avoids the risk of intrusion by these impurities and provides clear results with a low background level. - Disperse the beads well
If dispersion is insufficient following the FG beads washing process after the binding reaction with the proteins, there is the possibility of impurities remaining inside the bead clusters. Therefore it is necessary to disperse the beads well. Use the manual method to disperse the beads. (With ultrasound, there is the risk that the proteins will be damaged.)
Manual dispersion method
FAQ
Questions about FG beads
- Are the sizes of FG beads® all equal?
The sizes are 0.2 μm. The CV value indicating variation is approximately 30%.
- What is the linker length of FG beads®?
It is about 1 nm.
- What is the number of particles per 1 mg of FG beads®?
It is about 1.8×1011 particles.
- What types of the FG beads are there?
At present, 13 types of FG beads® are available, with different surface modificationssuch as functional groups or bound proteins. Additions to the lineup are currently being considered. In addition to FG beads®, there are also fluorescent beads.
- What is the approximate quantity of the functional groups on FG beads®?
The amount of epoxy on Plain beads is 1 μmol per 1 mg of beads. The amount of functional groups on COOH beads and NHS beads is 200 - 300 nmol per 1 mg of beads.
- Are the FG beads® resistant to heat?
There are no problems up to approximately 70°C. However the type of beads having high functional reactivity of the linker end may come under an influence by the heat.
- Are the FG beads® resistant to pH?
There are no problems within the range of pH2 - pH11. However the protein immobilized on beads may denature and the performance may decrease.
- Is there the resistance to organic solvents of FG beads® in addition to DMF?
Ligands can be immobilized on the surfaces of FG beads® in various organic solvents, such as methanol, DMF, DMSO, THF, ethyl acetate, pyridine, dioxane, toluene, dichloromethane, chloroform, etc.
- Can the FG beads® be reused?
It is believed that the beads can be reused when salt elution or drug elution was performed. However boil elution causes degeneration of the FG beads® and makes it impossible for them to be reused.
- How should one select a bead type?
Select the FG beads® with the optimal function group according to the substance or protein that you will bind, or select streptavidin beads
Questions about chemical pulldown and protein-protein interaction
- What is the general process used to bind ligands to the beads?
When binding ligands to NHS beads, the amount of ester content per unit weight of the beads is approximately 200 nmol/mg. For the binding reaction, 3 concentrations are prepared (ligand solutions: 0.1, 0.3, 1 mM) in order to determine the optimal concentration. The amount of bound ligands depends on the ligand properties, and varies approximately from several nmol to 100 nmol. An amount of several nmol to several tens of nmol is best for achieving purification results. An excessive amount of ligands results in steric hindrance and has a negative effect on affinity purification.
- What amount of the beads is required?
20 mg of beads are sufficient for 2 studies of the affinity purification conditions. A total of approximately 50 mg of beads are necessary in order to secure a certain amount of the purified target proteins or other substance after the study of the conditions.
- What are the important points when designing a ligand?
When designing a ligand in order to introduce a functional group for the parent compound of the protein you seek to identify, because the protein which will be purified varies depending on the location where the function group is introduced, it is recommended that receptor identification tests be performed by binding multiple ligands at different linker binding locations on the same parent compound. In the event that there is no functional group capable of binding to the ligand, it is necessary to change the chemical structure of the compound itself.
- Is it possible to bind secondary amines?
Secondary amines can be bound to NHS beads. However when both primary amine and secondary amine are present, the reaction will occur selectively with the primary amine.
- How are beads stored after the ligands are bound to them?
When the amount of bound ligands is large, the beads become hydrophobic and dispersibility decreases. Therefore the beads are stored in a 50% methanol solution, not in water.
- Are there any methods other than HPLC for verifying whether or not ligand binding has been successful?
The proteins can be actually bound to the ligands in order to verify binding. At a binding concentration of 0 mM, there is no protein binding. If the protein band increases as the binding concentration increases, then good binding has been verified.
- How strong is the affinity for the proteins that are affinity purified?
Ordinarily the affinity for the ligands of the proteins to be affinity purified is expressed as a dissociation constant (Kd) of 10-6 M or below. If the affinity is too low, there is the risk of a higher level of background noise.
- What is the purification efficiency?
When ligands with a relatively strong bonding strength are used, the results of studies by our company show a purification efficiency of approximately 50%. However the actual efficiency is highly dependent on the properties of the individual ligands and varies on a case-by-case basis.
- What is the optimal bead type for binding proteins?
The best beads are the NHS beads with activated COOH radicals on the COOH bead surface. The ε-NH2 radicals in the protein lysine residue bond with the COOH radicals on the bead surface. It is also possible to bind proteins to the epoxy beads.
- When binding proteins, what should be done if there is lysine residue at a location related to binding with the target substance?
It may be possible to bind the proteins by introducing a His-tag or by using cysteine residue sulfhydryl radicals (SH radicals).
- What is the efficiency when binding proteins?
If binding is performed when 50 mg of protein is supplied to 1 mg of NHS beads, approximately 10 – 50mg will be bound, although the result varies depending on the protein. It is possible to increase the amount that is bound by increasing the amount supplied.
- What is the optimal bead type for binding peptides?
The best beads are the NHS beads with activated COOH radicals on the COOH bead surface. The ε-NH2 radicals in the protein lysine residue bond with the COOH radicals on the bead surface. It is also possible to bind peptides to the epoxy beads.
- How is the cell extract prepared?
We recommend the Dignam method. However because the Dignam method requires a large quantity of cells, we recommend the NP-40 lysis method when preparing small quantities.
- Is there any problem with using frozen stock homogenate?
There is no problem. However perform centrifuge separation before using to remove any impurities (such as degenerated proteins).
- How much protein supply is necessary?
Serum proteins or extracts from cultivated cells or tissues can be used. When using cultivated cells as the protein source, 109 nuclear proteins or 107 - 109 cellular proteins are required.
- Can affinity purification be used with membrane proteins such as GPCRs and ion channels?
While it is not impossible, it is not easy to use affinity purification by this method for GPCRs and ion channels. There are cases when affinity purification can be used by solubilizing the membrane protein with a surfactant.
- Is this because the background level high?
It is necessary to consider the optimal conditions by varying the concentration of ligand binding to the FG beads®, the salt concentration in the buffer, and the surfactant concentration. This can sometimes be improved by carefully performing dispersion when screening. Insufficient masking is also a possibility, so it is necessary to suitably mask the functional groups which are not bound by the ligands.
- What should be done when a large number of bound protein bands are detected?
Change the buffer composition and detect a more highly specific band. Alternatively, it is necessary to perform competitive inhibition tests and drug elution, and verify whether or not the band is specific to that ligand.
- Is it necessary to use the recommended buffer as the binding buffer?
There are no problems with using TBS in place of HEPES, and NaCl in place of KCl.
- Why is it that both salt elution and boil elution are performed for elution?
This is because bonds are divided between somewhat weaker bonds (which can be broken by salt) and strong bonds (which can be broken by a sample buffer plus heating). However there is no problem with performing boil elution alone.
Questions about immunoprecipitation
-
Can I quantify the immobilization amount of the antibodies on the beads?
It can be carried out by direct quantification of the immobilized antibodies. please refer to Protocol 107 on our website.
- What is the optimal bead type for immobiliding antibodies?
The best beads are the NHS beads with activated COOH radicals on the COOH bead surface. The ε-NH2 radicals in the antibody lysine residue bond with the COOH radicals on the bead surface. It is also possible to bind antibodies to streptavidin and NeutrAvidinTM beads with biotin-labeled antibodies.
- What is the efficiency when immobiliding antibodies?
If binding is performed when 50 μg of antibodies is supplied to 1 mg of NHS beads, approximately 20 – 50 μg will be bound, although the result varies depending on the antibody animal, subclass, and clone. It is possible to increase the amount that is bound by increasing the amount on feed. When streptavidin beads are used, up to approximately 10 μg of biotin-modified antibodies will bond to 1 mg of streptavidin beads or NeutrAvidinTM beads, although this also varies depending on the antibody.
- Can I disperse antibody-immobilized beads by ultrasonic device?
Please perform the dispersion of the beads by the manual agitation. When you cannot disperse the beads easily, disperse them in a short time by using an ice-cold ultrasonic homogenizer or ultrasonic washer. Please refer to the video on website for details.
- When immobilizing antibodies to beads, the beads may adhere to the wall of the tube. Is there a way to suppress this?
It may be improved by stirring with a microtube mixer instead of inverting mixing. And please use a protein low bind tube.
- How can I improve dispersibility of antibodies immobilized beads?
Antibodies or proteins immobilized beads may easily precipitate. Please disperse the beads well before use. In addition, dispersibility improves if salt is removed from the buffer.
Questions about immobilization of DNA and RNA
-
What is the optimal bead type for binding DNA?
Single-strand and double-strand DNA can be immobilized to the Plain beads.
- What method is used to bind RNA?
Either bind synthesized RNA with a suitable functional group (amino group, SH radical, etc.) introduced onto the ends, or by binding DNA in advance and using hybridization to bind the RNA.
References
The Latest Reference Documents (Tamagawa Seiki Co., Ltd. web site)
- M, Watanabe et al., Oncogenesis, 6, e311 (2017)
- Y, Nishiya et al., Bioorg. Med. Chem. Lett., 27, 834 (2017)
- L. Jia et al., Biochem. Eng. J., 114, 268 (2016)
- T, Taniguchi et al., Cell Death Dis., 7, e2211 (2016)
- Jaiyam Sharma et al., Sensing and Bio-Sensing Research 9, 7?12 (2016)
- Y. kabe et al., Nature Communications, 7:11030 (2016)
- Y. Soeda et al., Nature Communications, 6:10216 (2015).
Downloads
- FG beads
 (PDF 1.9 MB)
(PDF 1.9 MB)
Ordering Information
FG beads
Others
(Storage) RT: Room temperature, A: Cool and dark, R: Refrigerator, F: Freeze